PseudotimeDE: inference of differential gene expression along cell pseudotime with well-calibrated p-values from single-cell RNA sequencing data | bioRxiv
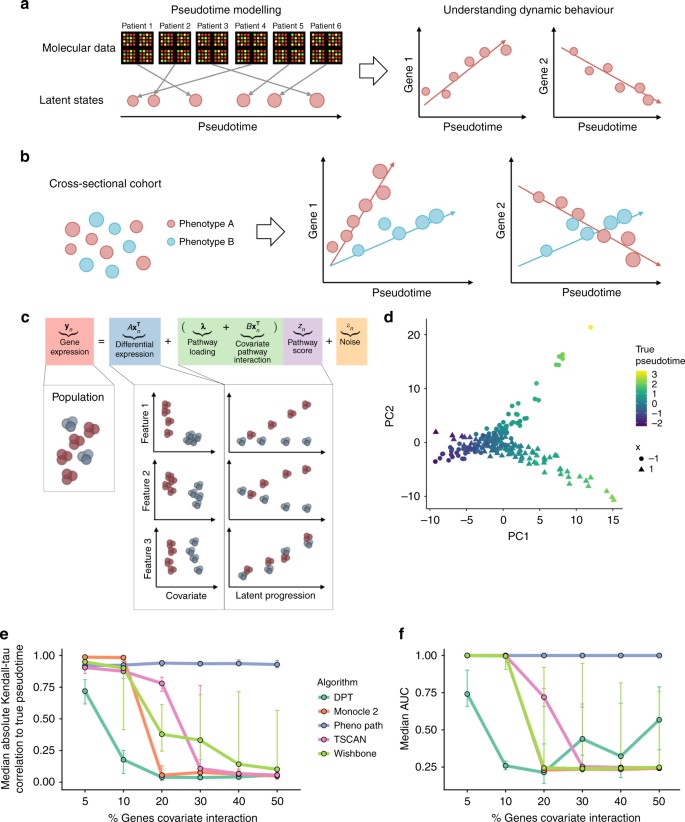
Uncovering pseudotemporal trajectories with covariates from single cell and bulk expression data | Nature Communications

JCI Insight - Single-cell analysis of fate-mapped macrophages reveals heterogeneity, including stem-like properties, during atherosclerosis progression and regression
Single-Cell Trajectory Analysis Using Monocle 2 and scGPS Pseudotime... | Download Scientific Diagram

Unsupervised Trajectory Analysis of Single-Cell RNA-Seq and Imaging Data Reveals Alternative Tuft Cell Origins in the Gut: Cell Systems

Spatiotemporal Developmental Trajectories in the Arabidopsis Root Revealed Using High-Throughput Single-Cell RNA Sequencing - ScienceDirect
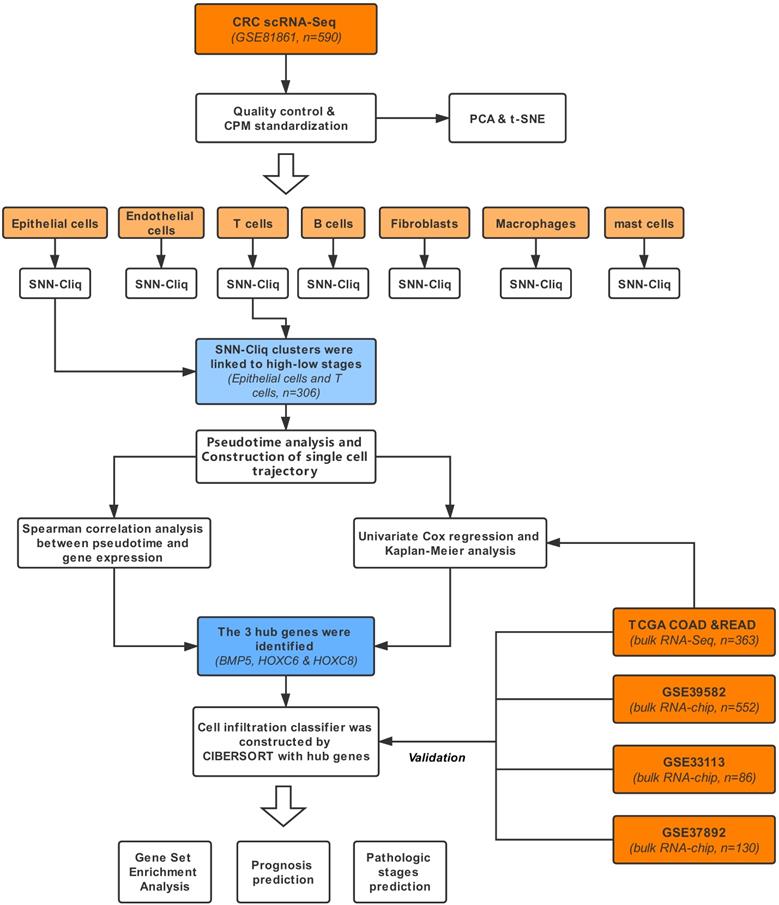
Pathologic evolution-related Gene Analysis based on both single-cell and bulk transcriptomics in Colorectal Cancer

PseudotimeDE, statistical method that outperforms existing methods in its detection power | UCLA Computational Medicine

Global Dynamic Molecular Profiles of Stomatal Lineage Cell Development by Single-Cell RNA Sequencing | bioRxiv
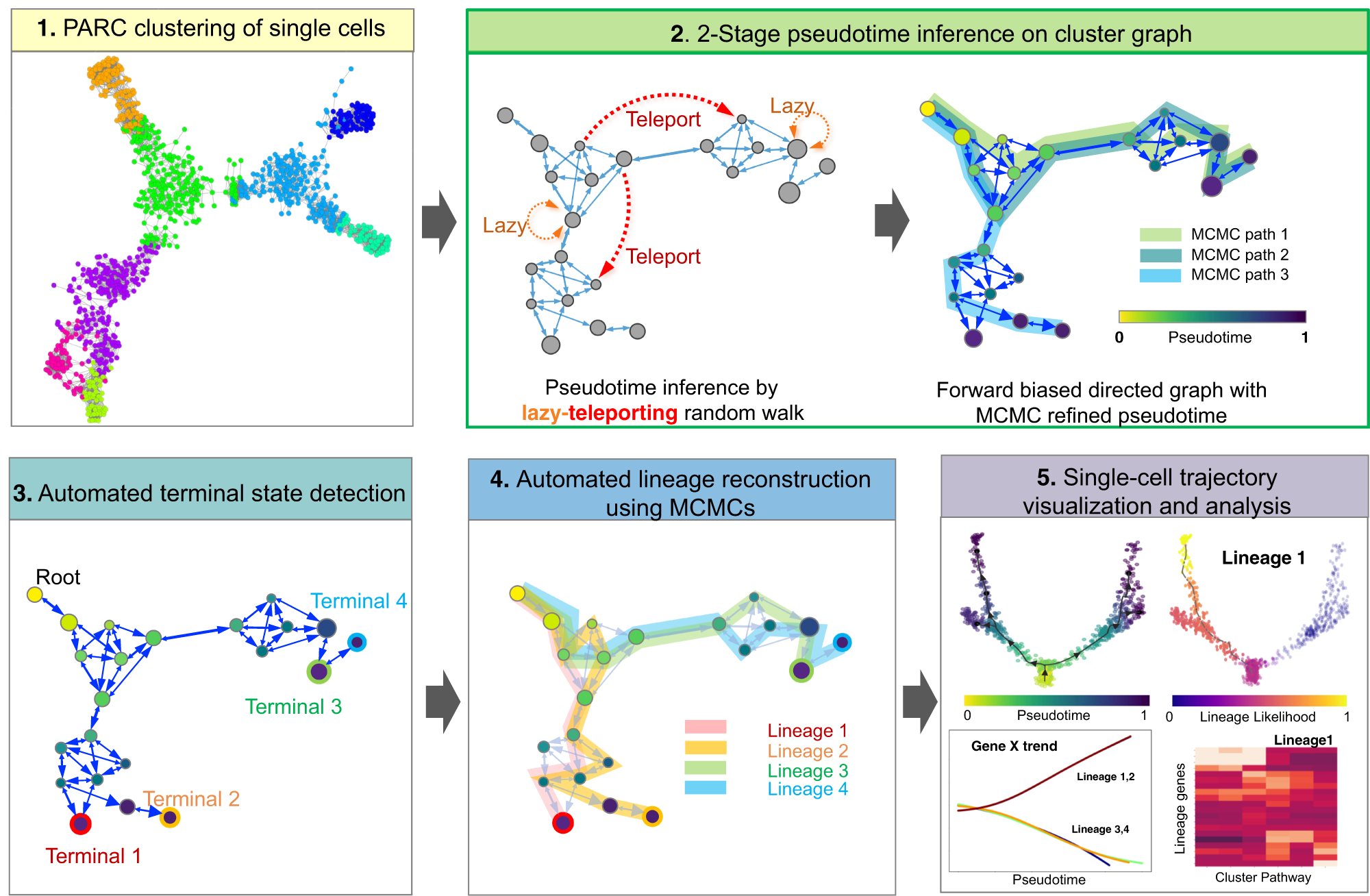
Generalized and scalable trajectory inference in single-cell omics data with VIA | Nature Communications
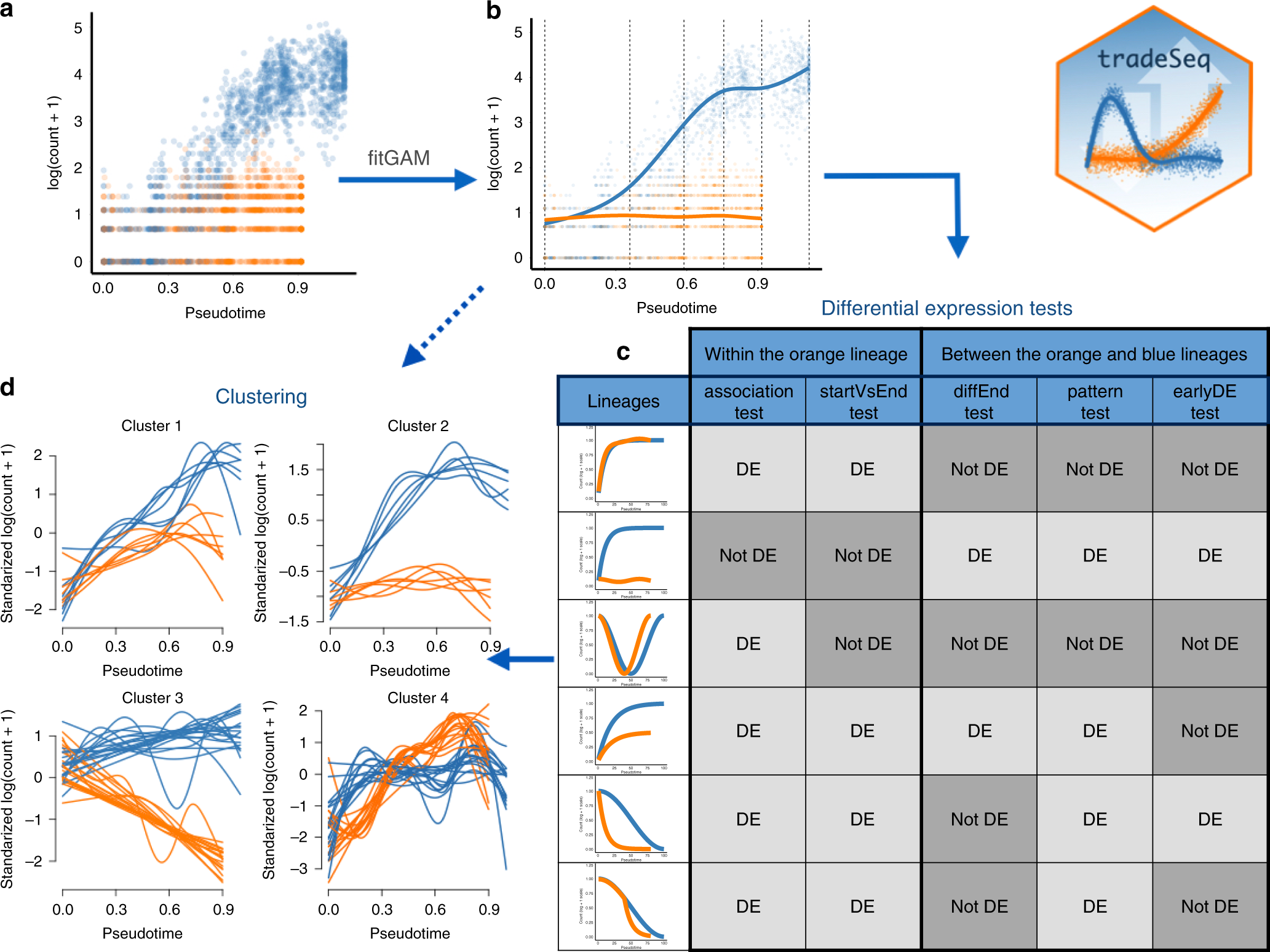
Trajectory-based differential expression analysis for single-cell sequencing data | Nature Communications